Summer 2010 — R: ggplot2 Intro
| <-General: Graphics | Home |
Contents
Intro
When it comes to producing graphics in R, there are basically three options for your average user.
- base graphics
lattice ggplot2
I've written up a pretty comprehensive description for use of base graphics here, and don't intend to extend beyond that. Base graphics are attractive, and flexible, but when it comes to creating more complex plots, like this one, the code to create it become more cumbersome.
Both
The website for ggplot2 is here: http://had.co.nz/ggplot2/. It I would highly suggest getting a copy of the manual: Amazon (as of July 2010, it looks like you can buy it new for cheaper than used!).
ggplot2 Basics
Plots convey information through various aspects of their aesthetics. Some aesthetics that plots use are:
- x position
- y position
- size of elements
- shape of elements
- color of elements
The elements in a plot are geometric shapes, like
- points
- lines
- line segments
- bars
- text
Some of these geometries have their own particular aesthetics. For instance:
- points
- point shape
- point size
- lines
- line type
- line weight
- bars
- y minimum
- y maximum
- fill color
- outline color
- text
- label value
There are other basics of these graphics that you can adjust, like the scaling of the aesthetics, and the positions of the geometries.
The values represented in the plot are the product of various statistics. If you just plot the raw data, you can think of each point representing the identity statistic. Many bar charts represent the mean or median statistic. Histograms are bar charts where the bars represent the binned count or density statistic.
Layer by Layer
There's a quick plotting function in
All
data - The data frame containing the data to be plotted
aes() - The aesthetic mappings to pass on to the plot elements
As you can see, the second argument,
The next step in creating a plot is to add one or more layers. Let's start with the an example from the
?mpg
summary(mpg)
p <- ggplot(mpg, aes(displ, hwy))
If you just type
p + geom_point()
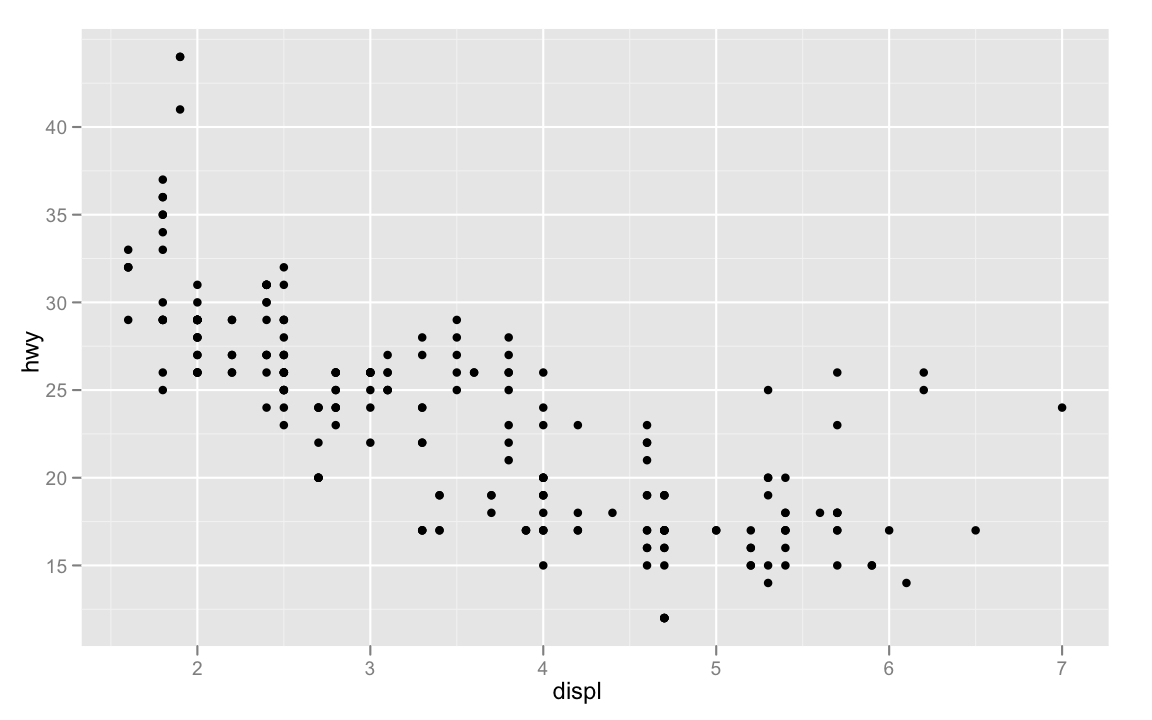
You add geometries to a plot with one of the
ggplot(mpg, aes(displ, hwy))+
geom_point()
Notice how we didn't pass any arguments to
The best way to demonstrate this is to make a few nonsensical plots. First, we'll create the same plot as above, but also connect all the points with a line.
ggplot(mpg, aes(displ, hwy))+
geom_point()+
geom_line()

Now, we're representing the x and y variables with points and a line, connecting all the points. This isn't a very meaningful plot for this data.
Next, we'll color the points according to the number of cylinders in the engine, treating number of cylinders as a nominal
variable. We'll pass this color mapping to
ggplot(mpg, aes(displ, hwy))+
geom_point(aes(color = factor(cyl)))+
geom_line()

The points are colored, the line is not, and a legend has automatically been added.
Next, we'll pass the color mapping to the line, not the points.
ggplot(mpg, aes(displ, hwy))+
geom_point()+
geom_line(aes(color = factor(cyl)))

Now the line is colored, and the points are not. It's kind of hard to tell with this plot, but lines which are different colors are not connected. The legend also represents the fact that lines are colored.
Finally, we can pass the color mapping to
ggplot(mpg, aes(displ, hwy, color = factor(cyl)))+
geom_point()+
geom_line()

Let's look at some other geoms with other data.
source("http://www.ling.upenn.edu/~joseff/rstudy/data/coins.R")
ggplot(coins, aes(coin, value))+
geom_point()

ggplot(coins, aes(coin, value))+
geom_bar()
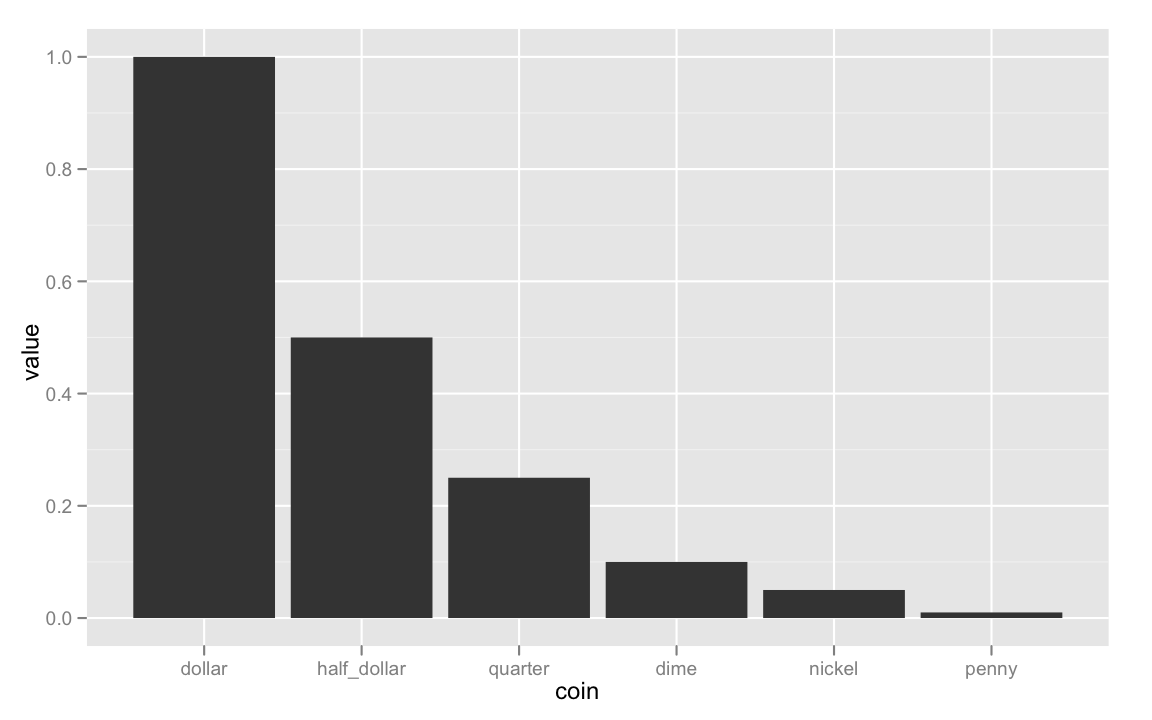
ggplot(coins, aes(coin, value * N))+
geom_point()

ggplot(coins, aes(coin, value * N))+
geom_bar()
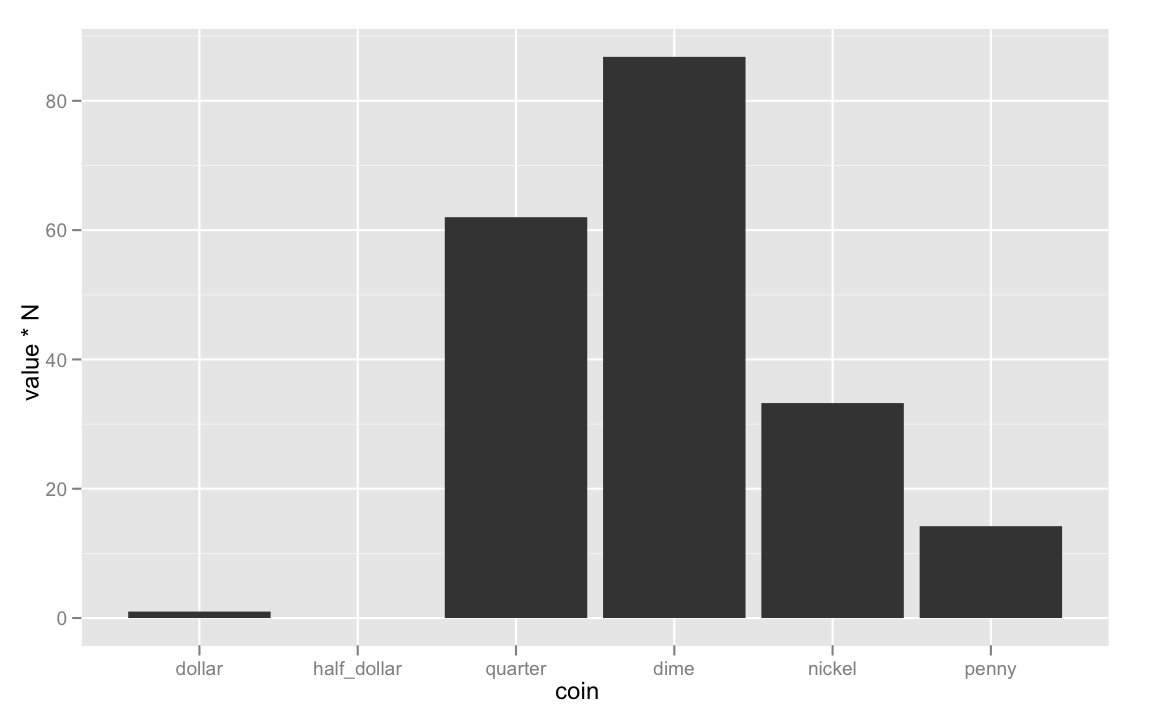
ggplot(coins, aes(coin, value * N))+
geom_bar(aes(color = coin))
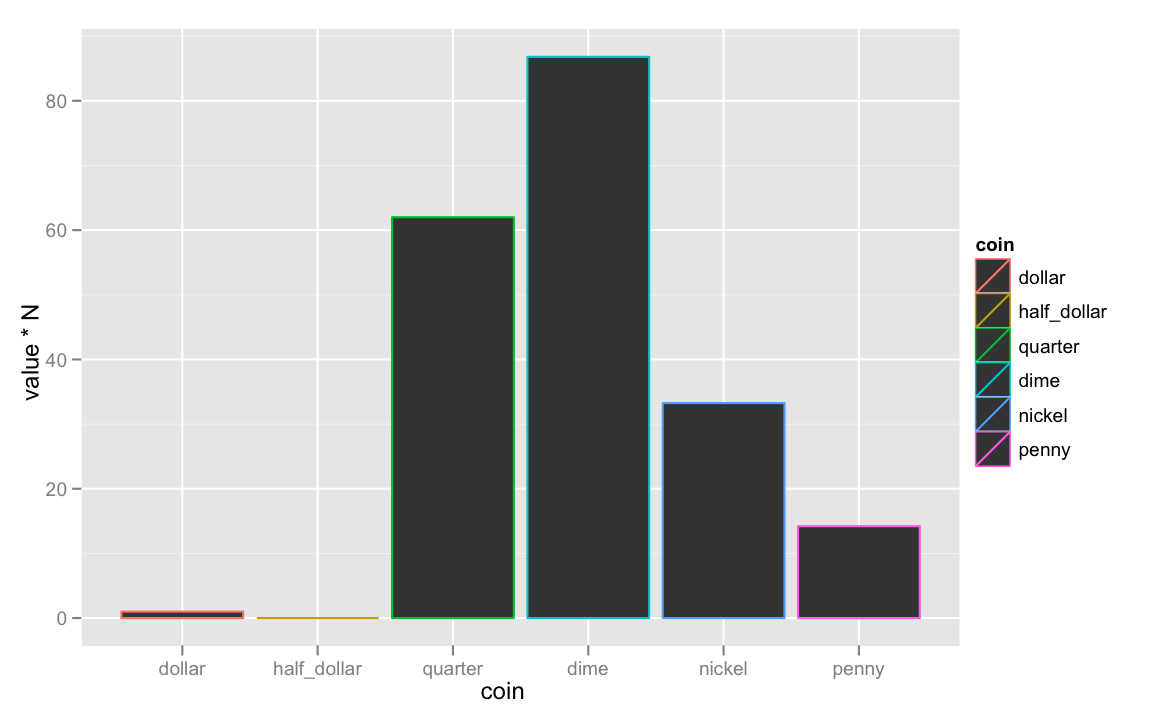
ggplot(coins, aes(coin, value * N))+
geom_bar(aes(fill = coin))

Displaying Statistics
You'll frequently want to add statistical analyses to your plots, or your plots may just be of statistical summaries anyway.
The most frequent statistic I use is a smoothing line with
p <- ggplot(mpg, aes(displ, hwy))
p + geom_point() + stat_smooth()

By default,
p + geom_point() + stat_smooth(method = "lm")
library(MASS)
p + geom_point() + stat_smooth(method = "rlm")
Now, statistics are represented with default geometries. For
p + stat_smooth(geom = "point")+stat_smooth(geom = "errorbar")

Geoms also have default statistics associated with them. For
## These should produce equivalent plots.
p + geom_point(stat = "smooth")
p + stat_smooth(geom = "point")
There exist some stats and geoms which have the same name. Adding either one to a plot will produce the same plot. Take
ggplot(mpg, aes(class, hwy))+
stat_boxplot()
#equivalent to
ggplot(mpg, aes(class, hwy))+
geom_boxplot()

The same actually goes for
#equivalent to
p + geom_smooth()
stat_summary()
One of the statistics,
fun.y - A function to produce
y aestheticss fun.ymax - A function to produce
ymax aesthetics fun.ymin - A function to produce
ymin aesthetics fun.data - A function to produce a named vector of aesthetics.
You pass a function to each of these arguments, and
ggplot(diamonds, aes(cut, price)) +
stat_summary(fun.y = median, geom = "bar")

median.quartile <- function(x){
out <- quantile(x, probs = c(0.25,0.5,0.75))
names(out) <- c("ymin","y","ymax")
return(out)
}
ggplot(diamonds, aes(cut, price)) +
stat_summary(fun.data = median.quartile, geom = "pointrange")

It's not necessary to write our own functions to plot quantile ranges or confidence intervals, however. There are a few summary functions
from the
mean_cl_normal() - Returns sample mean and 95% confidence intervals assuming normality.
mean_sdl() - Returns sample mean and a confidence interval based on the standard deviation times some constant
mean_cl_boot() - Uses a bootstrap method to determine a confidence interval for the sample mean without assuming normality.
median_hilow() - Returns the median and an upper and lower quantiles.
This code should produce the same pointrange plot as above.
ggplot(diamonds, aes(cut, price))+
stat_summary(fun.data = median_hilow, conf.int = 0.5)
We can add confidence intervals to a plot this way.
ggplot(mpg, aes(reorder(class, hwy, mean), hwy))+
stat_summary(fun.y = mean, geom = "bar")+
# stat_summary(fun.data = mean_cl_boot, geom = "errorbar")
# point ranges are prettier
stat_summary(fun.data = mean_cl_boot, geom = "pointrange")

You can also use
Grouping
Let's start by looking at how statistics are calculated by groups.
ggplot(mpg, aes(displ, hwy, color = factor(cyl)))+
geom_point()+
stat_smooth(method = "lm")

We mapped the
If we had decided to map the cylinder grouping to the point shape, rather than the point color, the statistic still would be computed over every subset.
ggplot(mpg, aes(displ, hwy, shape = factor(cyl)))+
geom_point()+
stat_smooth(method = "lm")

Now, the color of the smoothing lines aren't meaningful anymore, but they've been grouped exactly like we defined with the
We could also group by
ggplot(mpg, aes(displ, hwy, size = factor(cyl)))+
geom_point()+
stat_smooth(method = "lm")

There's a truly silly plot.
We could also define a grouping which is only meaningful for
ggplot(mpg, aes(displ, hwy, linetype = factor(cyl)))+
geom_point()+
stat_smooth(method = "lm")

If you use multiple grouping variables, groups will be defined as unique combinations of each of the levels.
ggplot(mpg, aes(displ, hwy, color = factor(cyl),
shape = factor(year),
linetype = factor(year)))+
geom_point()+
stat_smooth(method = "rlm")

Grouping isn't only useful for
ggplot(mpg, aes(class, hwy, fill = factor(year)))+
geom_boxplot()
#reorder class according to median(hwy)
ggplot(mpg, aes(reorder(class, hwy, median), hwy, fill = factor(year)))+
geom_boxplot()

Sometimes it will be necessary to properly define the groups in your data in order to plot it. Here's another example from the
library(nlme)
?Oxboys
ggplot(Oxboys, aes(age, height)) +
geom_point()

What if we wanted to draw a line for every subject? Simply adding
ggplot(Oxboys, aes(age, height)) +
geom_line()
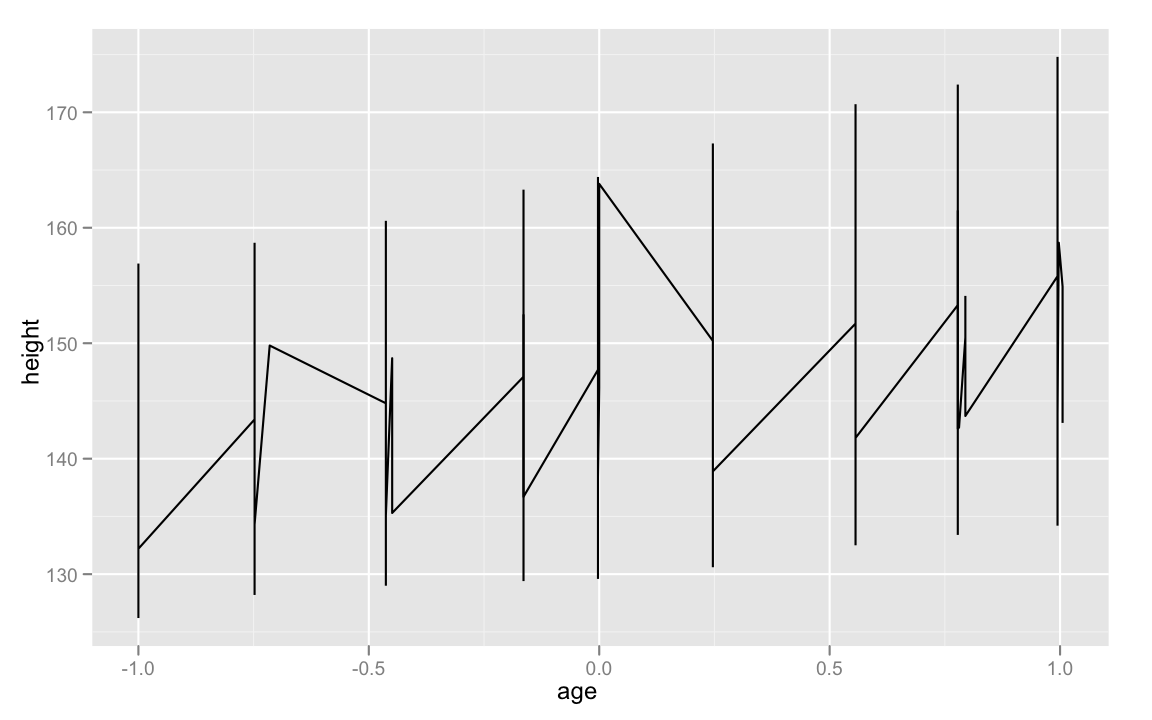
We need to define
So, what we'll do is define the
ggplot(Oxboys, aes(age, height, group = Subject)) +
geom_line()

Here's another example involving formant tracking.
jean <- read.csv("http://www.ling.upenn.edu/~joseff/data/jean2.csv")
ay <- subset(jean, VClass %in% c("ay","ay0"))
ay$VClass <- as.factor(as.character(ay$VClass))
ay.m <- melt(ay, id = c("Time","RTime", "Word","VClass"), measure = c("F1","F2"))
ggplot(ay.m, aes(RTime, value, color = VClass, linetype = variable)) +
geom_line()
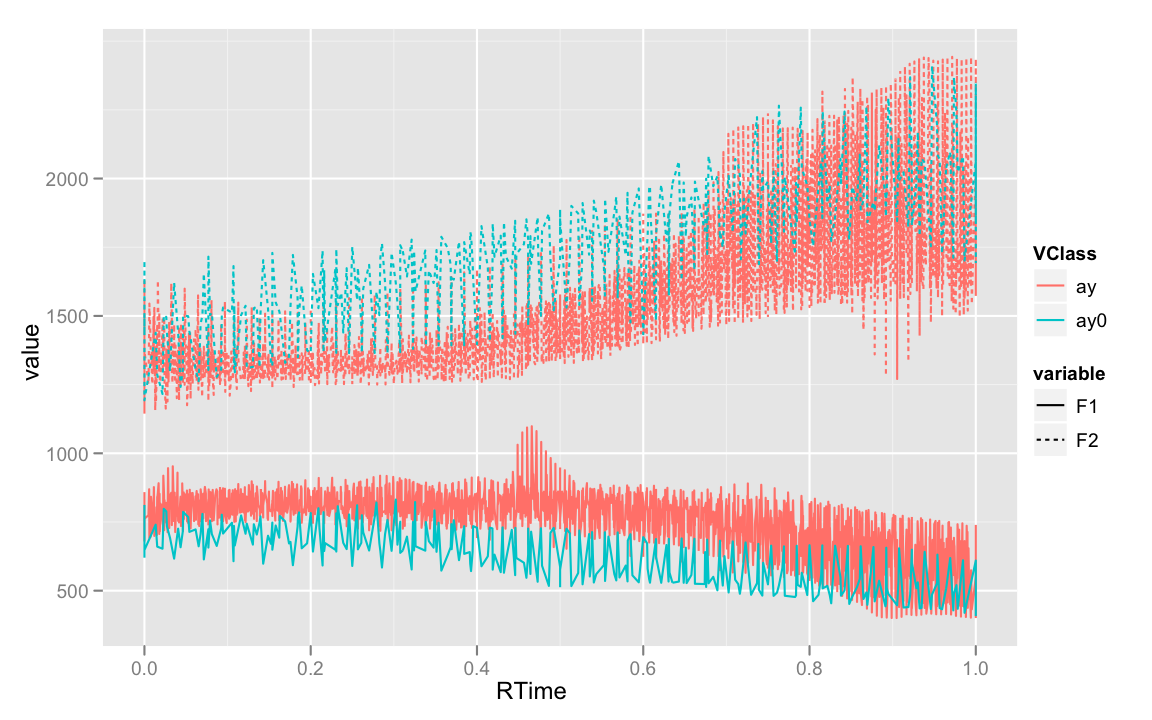
Clearly, this is wrong. We want to see lines for each word that was measured. Now, if we set
ggplot(ay.m, aes(RTime, value, color = VClass, group = Word)) +
geom_line()

This is also clearly awful. Taking into account that we want to plot two lines per word, we can define our groups using the interaction syntax.
ggplot(ay.m, aes(RTime, value, color = VClass, group = Word:variable)) +
geom_line()

F1 and F2 are pretty well separated, so it's probably not necessary to distinguish them with different linetypes.
If you ever want to draw connected lines over a nominal variable, you must define
The Philadelphia counties data we looked at last week is a good example.
source("http://www.ling.upenn.edu/~joseff/rstudy/plots/graphics/phila_bar.R")
Let's plot median income by educational attainmet.
ggplot(phil, aes(Level, value, color = Name, shape = Gender))+
geom_point()

The data's all displayed, but not very readable. Educational attainment has a clear and meaningful order, if not magnitude. We could add lines to this plot in a principled way. However, the following code should produce the same plot as above.
ggplot(phil, aes(Level, value, color = Name, shape = Gender))+
geom_point()+
geom_line()
Since
ggplot(phil, aes(Level, value, color = Name, shape = Gender, group = Name))+
geom_point()+
geom_line()

Again, not quite right, because we're plotting a line for every gender for every county.
#Changing shape = Gender to linetype = Gender
ggplot(phil, aes(Level, value,
color = Name,
linetype = Gender,
group = Gender:Name))+
geom_point()+
geom_line()

Positions
How geoms are positioned relative to eachother is another feature of plots that you might want to adjust. The possible position adjustments are
position_dodge() position_fill() position_identity() position_jitter() position_stack()
I'll demonstrate all of these except
## position = "stack"
## the default
ggplot(philcit, aes(Level, value, fill = Gender)) +
geom_bar(position = "stack")

ggplot(philcit, aes(Level, value, fill = Gender)) +
geom_bar(position = "dodge")

ggplot(philcit, aes(Level, value, fill = Gender)) +
geom_bar(position = "dodge")

ggplot(philcit, aes(Level, value, fill = Gender)) +
geom_bar(position = "identity", alpha = 0.3)

ggplot(ay, aes(F1, fill = VClass)) +
stat_density(aes(y = ..count..), position = "stack", color = "black")

ggplot(ay, aes(F1, fill = VClass)) +
stat_density(aes(y = ..count..), position = "fill", color = "black")

ggplot(ay, aes(F1, fill = VClass)) +
stat_density(aes(y = ..density..), position = "identity", color = "black", alpha = 0.5)

ggplot(jean, aes(-F2, reorder(VClass, -F2, mean)))+
geom_point(position = position_jitter(height = 0.25), alpha = 0.3)

ggplot(jean, aes(reorder(VClass, -F1, mean),-F1))+
geom_point(position = "jitter", alpha = 0.3)

You can also use jittered points as a kind of rug for plots of categorical data.
donner<-read.csv("http://www.ling.upenn.edu/~joseff/data/donner.csv")
ggplot(donner, aes(AGE, NFATE, color = GENDER))+
geom_point(position = position_jitter(height = 0.02, width = 0)) +
stat_smooth(method = "glm", family = binomial, formula = y ~ poly(x,2))

The jittered points are an ok built-in way to get this rug, but they're a little messy. I figured out a way to add cleaner points this way.
donner <- arrange(donner, GENDER)
donner <- ddply(donner, .(AGE, NFATE), transform, stack = (0:(length(AGE)-1))*0.015)
ggplot(donner, aes(AGE, NFATE, color = GENDER))+
geom_point(aes(y = abs(NFATE - stack))) +
stat_smooth(method = "glm", family = binomial, formula = y ~ poly(x,2))
Faceting
A very useful kind of visualization technique is the small multiple. In a small multiple visualization, you create many of the same
plot for multiple subsets of the data.
ggplot(mpg, aes(displ, hwy))+
geom_point()+
stat_smooth()+
facet_wrap(~year)

ggplot(mpg, aes(displ, hwy))+
geom_point()+
facet_wrap(~manufacturer)

Two things should be clear right off the bat. First, facets create further subsets for computing statistics over. Second, the x and y scales of each plot are the same in each facet. This is something that can be toggled, but doing so will usually eliminate the usefulness of creating a small multiple in the first place.
You can aslo facet by two variables using
?tips
ggplot(tips, aes(size, tip/total_bill))+
geom_point(position = position_jitter(width = 0.2, height = 0)) +
facet_grid(time ~ sex)

It looks like it was a male bill payer in a dinner party of two who tipped 70%. I'll leave all possible sociological analyses up to the reader.
Facet Scales
Usually you will want all of your facets to have the same x and y scales. If you're plotting the same data in each facet, having free
scales on each of the facets will ruin comparability across facets. However, sometimes it will be appropriate to have free scales. For instance,
when we plotted international data for men and women on a few different measures,
it was necessary to have free scales. I did this by passing
ggplot(data = gender.comp, aes(Male, Female))+
geom_abline(colour = "grey80")+
geom_point(alpha = 0.6)+
facet_wrap(~Measure, scales = "free")
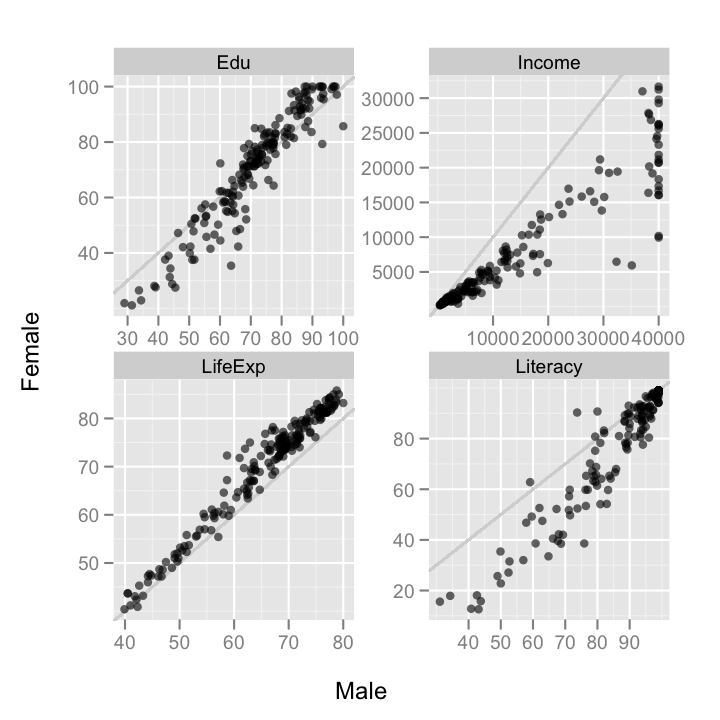
Income, LifeExpectancy, Literacy and Education are all measured on different scales with widely varying magnitude. If I hadn't
passed

The income scale completely overwhelms he others.
Sometimes, you'll only want one or the other scales to be free. To do this, pass
bp <- read.csv("http://github.com/ezgraphs/R-Programs/raw/master/BP_Oil_Recovery.csv")
bp$End.Period <- as.Date(bp$End.Period)
#Use with facet_wrap()
ggplot(bp, aes(End.Period, Recovery.Rate)) +
geom_area()+
facet_wrap(~Type, scales = "free_y")
#Doesn't work if you facet vertically with facet_grid() 
ggplot(bp, aes(End.Period, Recovery.Rate)) +
geom_area()+
facet_grid(.~Type, scales = "free_y")
#Facet horizontally 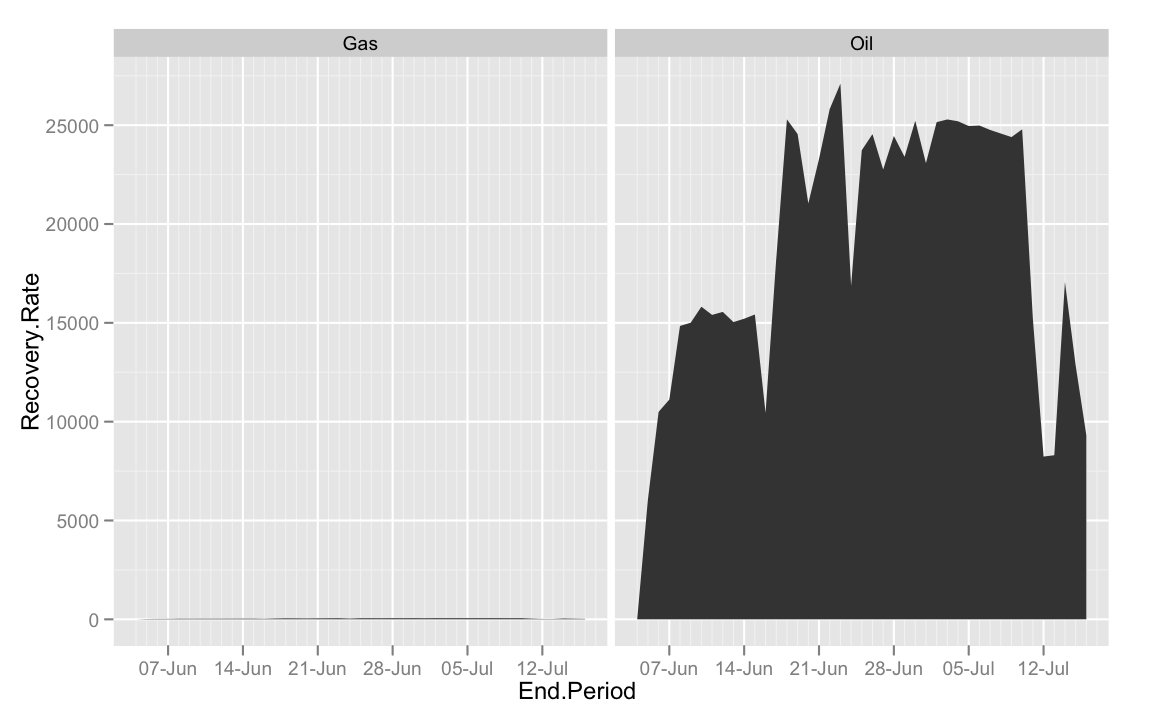
ggplot(bp, aes(End.Period, Recovery.Rate)) +
geom_area()+
facet_grid(Type~., scales = "free_y")

For those interested in these numbers, I'd suggest listening to this On The Media story about how the commonly reported volume of spilled oil in the Exxon Valdez disaster was possibly drastically underestimated.
Scales
Every aesthetic which is mapped to the data expresses the magnitude if its value along some scale. You can adjust these scales
using the
Almost all scales have a common set of arguments.
name - The text label for the scale
limits - The maximum and minimum values to be included in the scale
breaks - The labeled breaks for the data
labels - Labels for the breaks
trans - Transformation to use on the data.
The function calls for various scales are formatted like this:
x and y scales
The most common scale adjustments I do are for the
Here are some examples of identical scale manipulations.
p <- ggplot(mpg, aes(displ, hwy)) + geom_point()
p + scale_x_continuous(label = "Engine Displacement in Liters")
#or
p + xlab("Engine Displacement in Liters")
p + scale_x_continuous(limits = c(2,4))
#or
p + xlim(2, 4)
p + scale_x_continuous(trans = "log10)
#or
p + scale_x_log10()
An important thing to take into account is that adjustments to scales also transforms or throws away data for statistics. So for instance,
if you don't like that
color and fill Scales
The second most common scale adjustment I use is to the
Usually, you map either continuous or discrete data to colors in a plot. The default scale for discrete data is
p + scale_color_hue(label = "Cylinders")
Some people don't like the default discrete colors. With
library(RColorBrewer)
display.brewer.all()
I personally like
p + scale_color_brewer(pal = "Set1")
However, for this data we should probably consider one of the sequential palletes. The number of cylinders in an engine is an ordered variable after all.
p + scale_color_brewer(pal = "Blues")
p + scale_color_brewer(pal = "OrRd")
The range of possibilities with continuous color variables is huge. The default continuous color scale is
p <- ggplot(diamonds, aes(carat, price, fill = )) +
stat_density2d(aes(fill = ..density..), contour = F, geom = "tile") +
scale_x_log2()+
scale_y_log10()
p + scale_fill_gradient(high = "black", low = "white")